This website illustrates my academic research during my postdoctoral work with Harel Weinstein at Weill Cornell Medical College and PhD work with Vincent Voelz at Temple university.
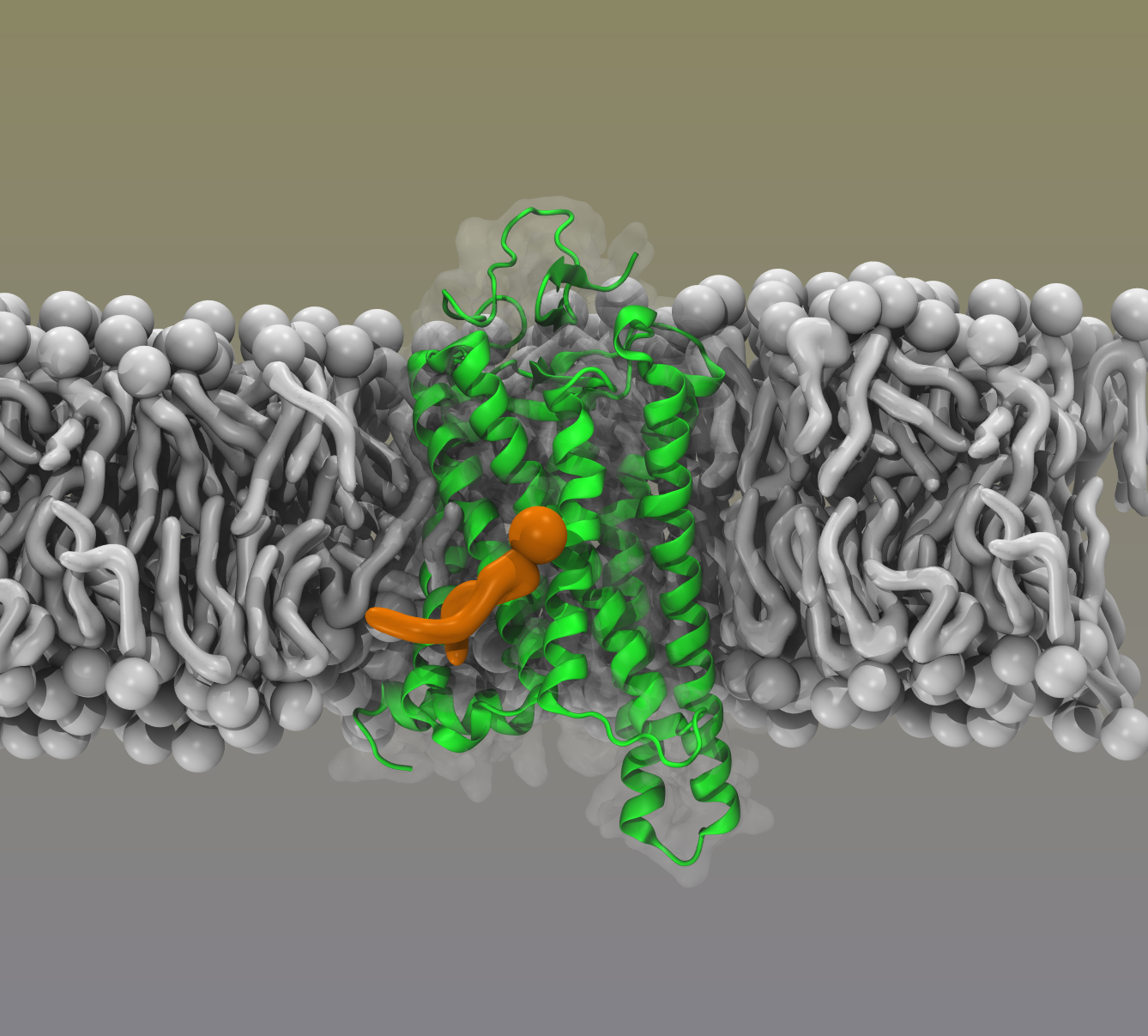
My postdoctoral academic research was focused on understanding function of membrane proteins that clear neurotransmitters from synaptic cleft. These family of proteins are called neurotransmitter:sodium symporters (NSS) since their function depends on electrochemical gradient of sodium ions across membrane. Important members of NSS family include dopamine transporter (DAT), serotonin transporter (SERT), and norepinephrine transporter (NTE) that remove dopamine, serotonin, and norepinephrine from synaptic cleft. Proper control of the amount of neurotransmitter in the synapse is important for many brain functions and imbalances are linked to brain related diseases like depression, bipolar disorders, ADHD, Schizophrenia, and Parkinson’s. In my research, I combine MD simulations with kinetic models based on Markov State Models (MSMs) to understand function of NSS family of proteins.
curriculum-vitae (PDF)