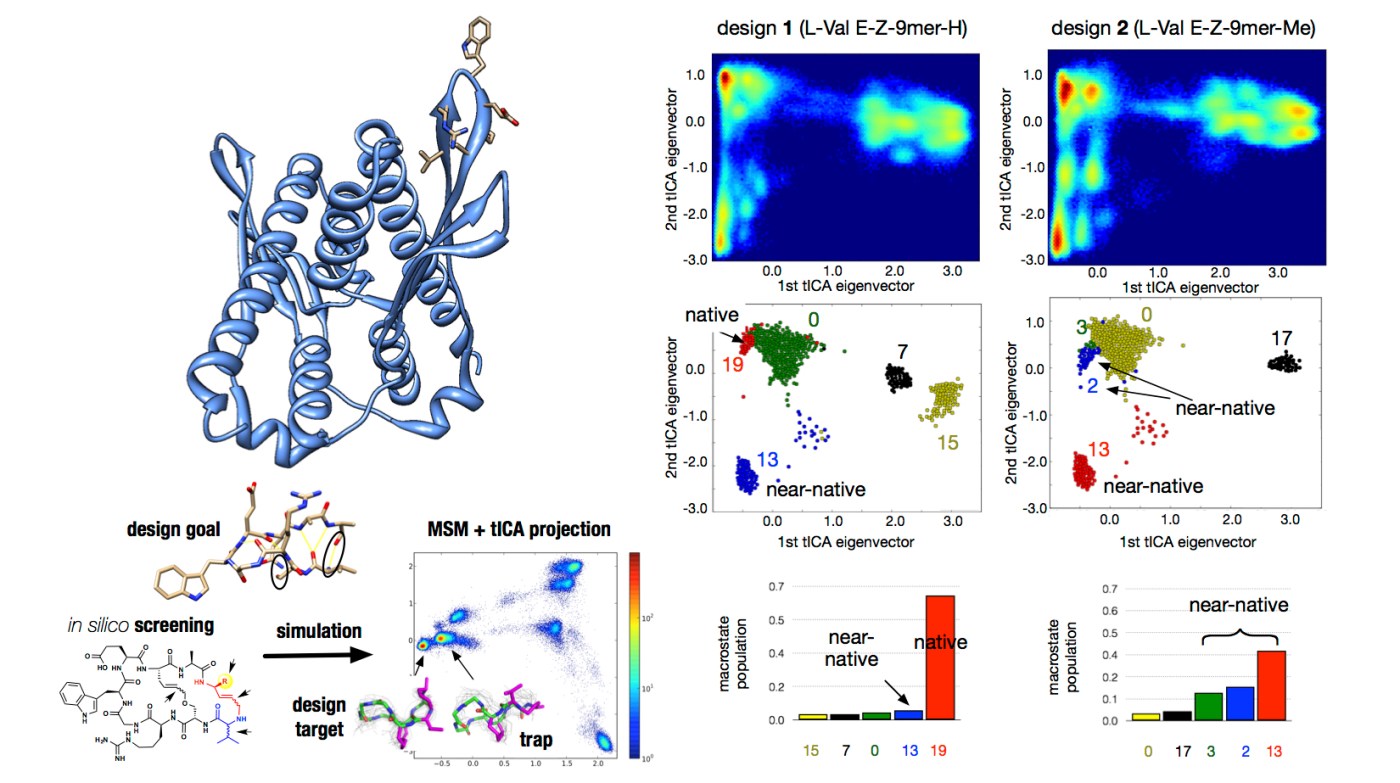
The ability of biofilms to attach surfaces is crucial for their life cycle. LapA is a protein that is responsible for this strong adhesion. Another protein called LapG can cleave N-terminal of LapA, hence freeing bacteria from surface. Normally, LapG is in interaction with LapD protein and therefore it is not free to cleave LapA. In this computational/experimental project, our goal is to design small peptidomemtics that can interrupt LapD-LapG interactions, therefore LapG will be free to cleave LapA. Initially, we designed peptidomimetics with 7 and 9 residues (7-mers and 9-mers), however, after fast computational screening using replica exchange implicit solvent MD, 9-mer designs seen to be more potent.
- Journal article: Computational screening and selection of cyclic peptide hairpin mimetics by molecular simulation and kinetic network models, Asghar M. Razavi, William W. Wuest, Vincent A. Voelz, Journal of Chemical Information and Modeling 54: 1425–1432, 2014
- Poster: Razavi, A. M., Wuest, W. M., & Voelz, V. A. (2014, August). Computational screening and selection of cyclic peptide hairpin mimetics by molecular simulation and kinetic network models. Poster presented at the ACS National Meeting, San Francisco, CA. [PDF]
About the author asgharrazavi
Asghar Razavi is a postdoctoral associate at the Department of Physiology and Biophysics at Weill Cornell Medical College of Cornell University. He received his Ph.D. in Computational Chemistry and Biophysics from Temple University, Philadelphia, USA. His current research at the Weinstein lab focuses on developing molecular level quantitative kinetic models to understand thermodynamics, kinetics, and conformational pathways during function of neurotransmitter transporters and G protein-coupled receptors.
-

June 23, 2017
-
June 23, 2017
-

January 7, 2017
All posts by asgharrazavi →About me
Dopamine Transporter